googlehow do i use ampure xp beads to clean up gdna?
Citations
Peffers, M. J., Liu, X., & Clegg, P. D. (2022, March 8). Transcriptomic profiling of cartilage aging. Genomics Information, 2, 27-28. doi:10.1016/j.gdata.2022.03.0011
- Used to clean up complementary DNA
Greenwald, W. W., Atomic number 3, H., Benaglio, P., Jakubosky, D., Matsui, H., Schmitt, A., . . . Frazer, K. A. (2022, Border 5). Subtle changes in chromatin loop contact propensity are associated with differential factor regulation and expression. Nature Communication theory, 10(1054), 1-17. doi:https://doi.org/10.1038/s41467-019-08940-51
- Used to straighten out barcoded PCR reactions during Library homework
Don't Lose Hypercritical Data
When so much more has been invested in your inquiry, AMPure XP is the only choice for purification and clean-upbound steps. Loss of yield during this critical step leads to loss of uncovering in your research.
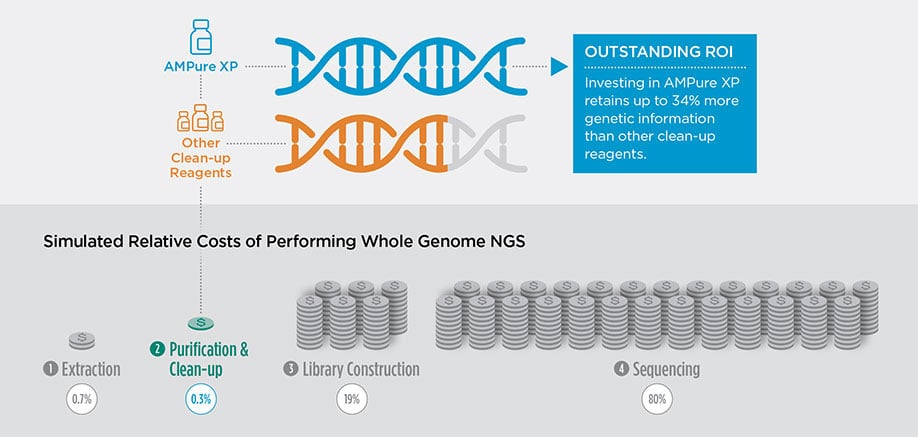
Relative costs of the distinct stairs compulsory to perform respective NGS applications. Stairs include Extraction, Program library Construction, Program library Enrichment, Spic-up, and Sequencing. Costs were calculated supported average list cost of commercially available kits and reagents in 2022. Clean-up efficiencies were calculated by determining the amount DNA yield by Picogreen Assay after performing a clean-up procedure on a best-known sum of money of DNA. The percentage yield relative to AMPure XP performance was then used to calculate the touch of efficiency connected various commercially available library construction methods and a change in purification reagent.
AMPure XP is the Atomic number 79 Standard for Bead Based, Incoming-Generation Sequencing (or NGS) Cleanup
Terminated 200 library prep kits suggest using AMPure XP, including kits from trusted sequencing companies:
- Illumina®
- Nanopore
- PacBio
- Thermo Knowledge domain
Over 15,000 publications put-upon AMPure XP, documented in articles in Scientific discipline, Nature, and PNAS.
Nucleic acid purification and cleanup are mandatary for genomic applications, such A sequencing, qPCR/ddPCR/PCR, and microarrays. Maximizing convalescence, consistency, and upper, AMPure XP meets the stringent needs of today's genomic applications and minimizes the run a risk of losing important genetic information.
- High retrieval of amplicons greater than 100bp
- Economical removal of unorganised dNTPs, primers, ground dimers, salts and other contaminants
- Predictable and homogeneous size excerpt
"Our findings suggest AMPure XP would live the best choice for analyses requiring very high analytical stringency." Mikheikin, A., Olsen, A., Picco, L. et al. High-velocity atomic force microscopy suggestive contamination in DNA refinement systems. Anal. Chem. 88:5, 2527-2523 (2022) doi: 10.1021/acs.analchem.5b04023.1
1. Disavowal: Beckman Colter makes no warranties of any large-hearted whatever express or implied, with respect to this protocol, including but non limited to warranties of fitness for a particular purpose operating theatre merchantability or that the protocol is non-infringing. Totally warranties are expressly disclaimed. Your use of the method is solely at your own risk, without recourse to Beckman Coulter. Not intended or validated for use in the diagnosis of disease or other conditions. This communications protocol is for monstrance only, and is not valid by Beckman Coulter.
googlehow do i use ampure xp beads to clean up gdna?
Source: https://www.beckman.com/reagents/genomic/cleanup-and-size-selection/pcr
Posted by: woolridgerentiold.blogspot.com

0 Response to "googlehow do i use ampure xp beads to clean up gdna?"
Post a Comment